AAV Vectors in Gene Therapy: Measuring & Optimizing Quality
Adeno-associated viruses (AAV) are promising vectors for in vivo gene therapy. They can deliver a genetic payload of up to 4.7 kb and infect a wide range of cell types with virtually no pathogenicity. AAV production is a stepwise process that begins with construction of a transfer plasmid and ends with packaging of the recombinant genome into viral particles (Figure 1). Generating preparations of AAV particles with high-fidelity DNA can be difficult due to inherent features of the AAV genome. Here, we present highlights from a recent AAV workshop to help you overcome common roadblocks and optimize the quality of your AAV vectors.
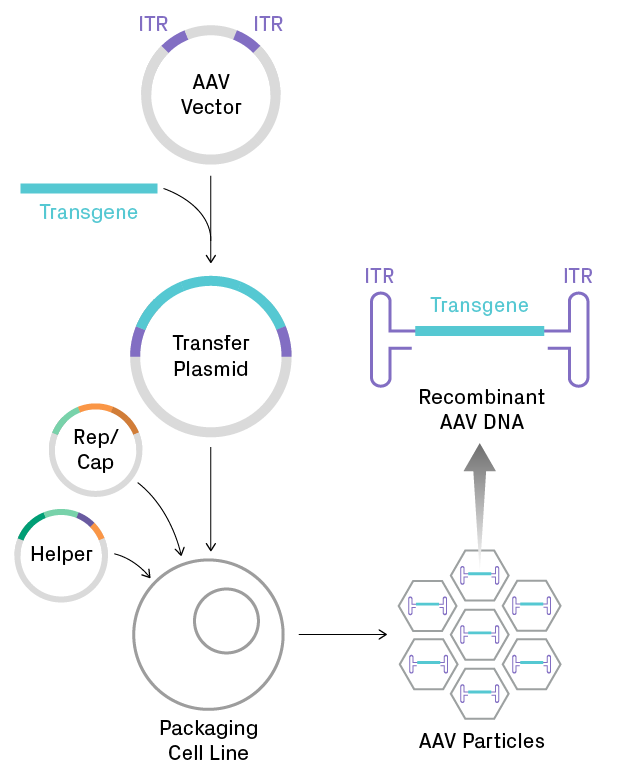
Figure 1. Production of recombinant AAV for gene therapy. A gene of interest is cloned into an AAV vector to create a transfer plasmid, which is then introduced into a eukaryotic cell line. Generating functional viral particles requires co-transfection with several viral genes, supplied by the Rep/Cap and helper plasmids. Recombinant single-stranded DNA, containing the transgene flanked by two inverted terminal repeat (ITR) sequences, is packaged into capsids. The ITRs form very stable hairpins, which are problematic for propagation and sequencing of AAV plasmids.
Validating the ITR Regions of AAV Vectors
The inverted terminal repeat (ITR) regions of AAV constructs can be very troublesome for cloning. These structures, usually 145 bases each, flank the transgene and form large hairpins in the single-stranded AAV genome (Figure 1). Due to the GC-rich, palindromic nature of ITRs, AAV plasmids are prone to spontaneous mutagenesis. Truncations frequently arise in the ITR regions when plasmids are propagated in bacteria.
Vigilance is key when working with AAV plasmids. Finding and addressing mutations as soon as possible will save considerable time and resources. Failure to spot an ITR truncation before packaging can severely compromise downstream experiments. ITR integrity should be checked in the AAV vector prior to cloning, especially if using a plasmid stock from a third party. It should also be verified after introduction of the transgene in the final preparation of the transfer plasmid since truncations can occur during scale-up of the DNA.
Sequence validation of AAV plasmids requires a specialized assay. Sanger sequencing is the gold standard for verifying plasmid sequences. However, the extensive secondary structure of ITR regions renders standard Sanger sequencing ineffective: chromatograms show a major drop in signal intensity near the start of the ITR sequence. Even enhanced Sanger protocols for difficult DNA templates often fail. AAV-ITR Sanger sequencing, developed specifically for AAV plasmids by Azenta Life Sciences, provides the most reliable method of ITR validation. It can read through entire ITR sequences, enabling you to confirm the integrity and clonality of your constructs.
Generating High-Fidelity AAV Plasmid Preparations
As mentioned above, ITR sequences are highly unstable. Complete or partial loss of an ITR sequence can confer a growth advantage to bacteria in culture, enabling de novo mutations to rapidly increase in frequency. As a result, generating plasmid preparations for AAV constructs with fully intact and clonal ITR regions is often a major challenge.
Two measures can be taken to optimize AAV plasmid prep. The first is to use a sufficiently robust and sensitive assay to detect mutations after DNA preparation, that is AAV-ITR Sanger sequencing. Without this, you cannot be confident about the fidelity of your plasmid prep. The second measure is to use a bacterial strain that maintains ITR integrity. Recombination-deficient strains will help reduce the frequency of mutagenesis, but strains commonly used in molecular cloning are still not sufficient for many AAV constructs. Again, a purpose-built solution provides the best option. Azenta Life Sciences developed E. coli strains optimized for ITR integrity. They significantly increase the stability of ITR sequences, outperforming commercial strains.
When mutations are detected in a plasmid stock, corrective actions are necessary. Preps with clonal truncated ITRs require replacement of the affected region, usually with a newly synthesized DNA fragment. For a mixed population of plasmids where some contain the truncated ITR, clones should be reisolated, screened using AAV-ITR sequencing, and amplified in a bacterial strain optimized for ITR stability.
For researchers interested in scaling up or constructing AAV plasmids using best current practices, we recommend AAV plasmid preparation and AAV plasmid synthesis solutions from Azenta Life Sciences (Figure 2). Both services rely on AAV-ITR sequencing for quality control before and after scale-up, employ proprietary E. coli strains to boost ITR stability, and offer ITR correction strategies for mutations. AAV plasmid synthesis also includes transgene synthesis and cloning for a convenient, end-to-end solution.
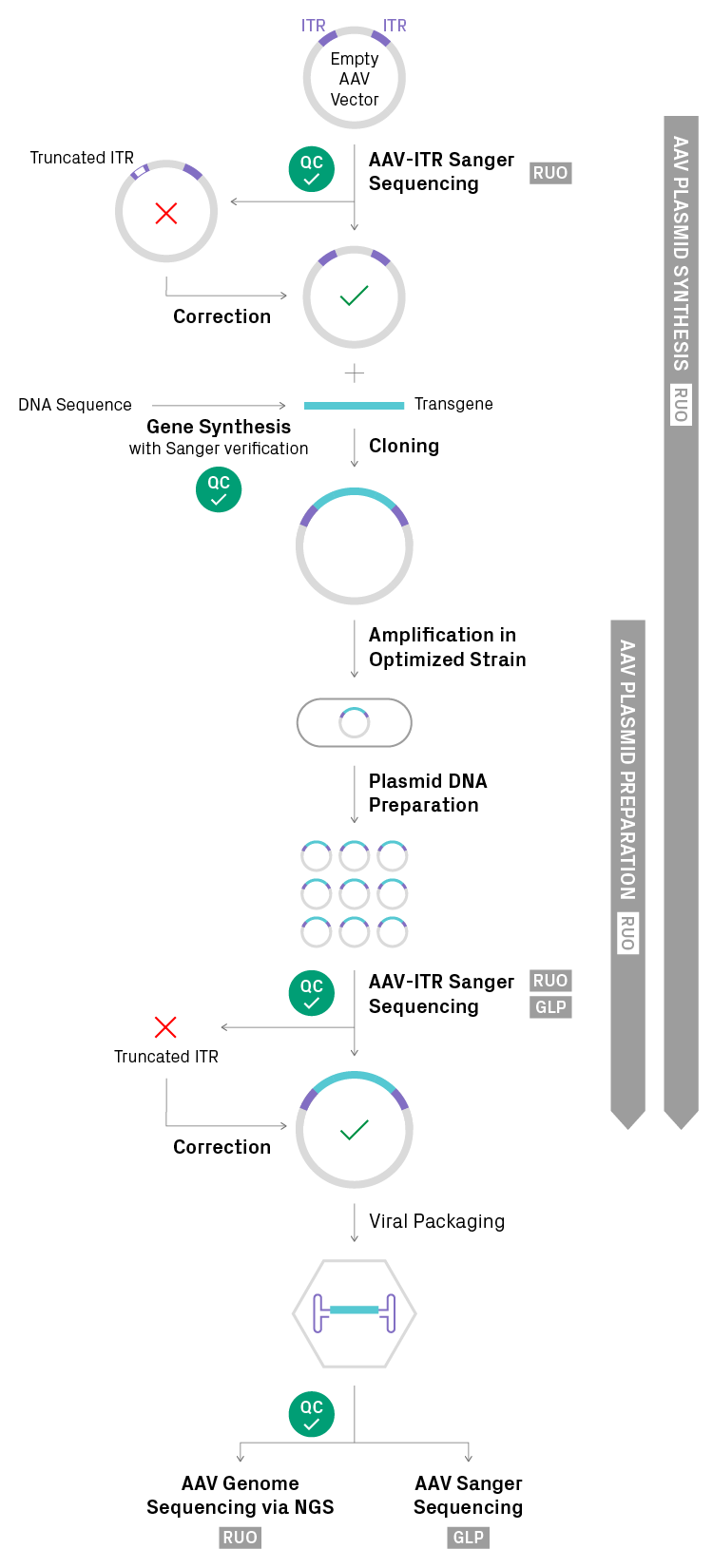
Figure 2. Optimized workflow for AAV construct assembly and validation. Sequencing-based quality control (QC) is performed at multiple stages of the process. The original stock of AAV plasmid is analyzed by AAV-ITR Sanger sequencing from Azenta Life Sciences to determine whether the population is clonal (i.e. homogeneous) and both ITR regions are fully intact. If mutations are detected after initial QC, corrective actions are available. The transgene is synthesized, sequence verified, and cloned into the AAV vector. The resulting plasmid is replicated in a proprietary E. coli strain that is optimized for ITR stability. The transfer plasmid preparation is validated by AAV-ITR sequencing and then introduced into a packaging cell line. The packaged DNA product is analyzed by next generation sequencing (NGS) or AAV-ITR sequencing. Azenta Life Sciences offers parts of the workflow at two regulatory levels: Research Use Only (RUO) and Good Laboratory Practice (GLP).
Verifying the DNA Packaged in AAV Particles
Once the transfer plasmid is constructed and verified, it’s transfected into a packaging cell line along with helper and Rep/Cap plasmids. The objective at this stage is to insert copies of the single-stranded recombinant AAV genome into functional viral particles. However, contaminant DNA and mutated sequences may be packaged as well, which can impact safety and/or efficacy of the gene therapy. Contaminant sequences can originate from the transfer plasmid backbone, Rep/Cap plasmid, helper plasmid, or host cell.
Analysis of the packaged product is thus necessary for quality control. Next generation sequencing (NGS) is the best way to measure the purity, clonality, and fidelity of the packaged DNA. Short- and long-read sequencing approaches can be used; each provides valuable information that is complementary to the other. Both approaches are recommended to obtain a full picture of the packaged product and can be performed in parallel by splitting the input sample.
With Illumina® short-read sequencing, the packaged DNA product is fragmented and amplified as part of the library preparation. Very deep sequencing enables highly sensitive and accurate quantification of all the DNA species in the sample. Once the Illumina data is mapped to all potential sources of DNA, you can measure the fraction of reads that represent recombinant AAV genome, the identity and frequency of contaminants, and the abundance of mutations in the AAV genome.
PacBio® long-read sequencing can provide similar information, albeit at a lower limit of detection since the DNA product is not amplified during library preparation and the sequencing depth is lower. The major advantage of PacBio sequencing is the ability to read full-length contiguous sequences of the DNA products. You can see exactly what molecules are getting packaged into particles. This method enables you to identify truncation hotspots, resolve the size of contaminants, and detect chimeras.
Validating AAV in Preclinical and Clinical Studies
As gene therapies move into the preclinical and clinical stages, rigor and reproducibility are essential. Safety and efficacy studies in animals, and eventually patients, require strict regulatory compliance and documentation, such as Good Laboratory Practice (GLP) for preclinical work. AAV-based gene therapies and the components used to make the final product should be thoroughly assayed for quality. This includes sequence confirmation of the AAV transfer plasmid, the packaging cell line, and the packaged viral DNA. GLP-compliant AAV sequencing is recommended for validating full AAV sequences, including the ITR regions, when preparing data for FDA submissions - such as IND, NDA, and BLA applications.
Conclusion
The ITR sequences of recombinant AAV genomes are necessary for viral function but pose significant challenges in the laboratory. Multiple quality control steps throughout the process are necessary to ensure that AAV constructs do not acquire mutations that can affect downstream safety or efficacy. Recent technological developments by Azenta Life Sciences enable reliable sequencing of the entire AAV construct as well as high-fidelity synthesis of the transfer plasmid.
To learn more, watch the recording of our AAV workshop and roundtable discussion, entitled “A Comprehensive Guide to Using Adeno-Associated Virus (AAV) Vectors in Gene Therapy”.