Fine-Tuning mRNA Structure for Better mRNA Therapies
The structure of mRNA has been known for decades, but only in recent years have researchers unlocked its potential for therapeutic development. With the right delivery vehicle, mRNA products can replace defective proteins in the cell, generate antigens for immunization (e.g., COVID vaccines), or edit the genome via CRISPR technology. Let’s review the structural features of a functional mRNA molecule and discuss how to optimize these for therapeutic applications.
mRNA Structure
For mRNA therapies, the objective is robust expression of the encoded protein. Transcripts that are efficiently translated and degraded slowly fare the best. Going from 5’ to 3’, components of an mRNA molecule include the 5’ cap, 5’ untranslated region (UTR), coding sequence, 3’ UTR, and poly(A) tail (Figure 1). Collectively they influence the amount of protein synthesized by dictating the transcript’s stability and translational efficiency in the cytosol. In the next few sections, we’ll examine each element.
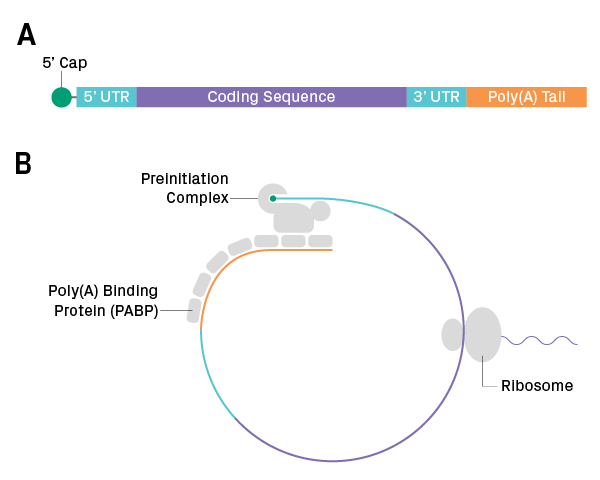
Figure 1. Structure of mRNA. (A) Anatomy of a eukaryotic transcript. (B) In the cytosol, the 5’ end of the transcript binds to the preinitiation complex, which recruits the ribosome to start translation. Poly(A) binding proteins at the 3’ end interact with the preinitiation complex, forming a loop structure.
5’ cap
Description: A modified guanosine (7-methylguanylate, abbreviated as m7G) attached to the 5’ end of the RNA molecule via a 5’ to 5’ triphosphate linkage.
Function in the cytosol: The 5’ cap binds to the preinitiation complex to promote translation. Also, its unusual attachment to the mRNA prevents degradation by 5’ to 3’ exonucleases.
Considerations for therapies: Nucleotide modifications adjacent to the 5’ cap are common in mammals. Methylation of the first nucleotide after the 5’ to 5’ triphosphate linkage, represented as m7GpppNm, creates a structure called cap 1. In contrast, mRNA with cap 0 (m7Gppp) lack this modification. Cap 1 is important for differentiating self from non-self RNA and thus evading the innate immune response1. Nucleotide analogues can be used during in vitro transcription to create cap 1 structures.
5’ Untranslated Region (5’ UTR)
Description: Nucleotide sequence upstream of start codon, excluding the 5’ cap.
Function in the cytosol: The 5’ UTR can regulate initiation of translation by interacting with the preinitiation complex or RNA-binding proteins. Stable secondary structures in the 5’ UTR, such as hairpins, can inhibit ribosome binding or scanning, lowering protein expression2.
Considerations for therapies: UTR sequences from highly expressed genes are typically used for mRNA therapies. The most popular are derived from the alpha and beta globin genes3. Adding an internal ribosomal entry site (IRES) to the 5’ UTR has been shown to boost translation efficiency. It can recruit the ribosome independently of the 5’ cap.
Coding Sequence (CDS)
Description: Sequence translated by the ribosomes, beginning with a start codon, and ending with a stop codon. Also known as the open reading frame (ORF).
Function in the cytosol: The CDS encodes the amino acid sequence of the protein.
Considerations for therapies: Codon optimization of the CDS can enhance protein expression. Due to redundancy in the genetic code, multiple codons can specify the same amino acid. However, synonymous codes are typically used at unequal frequencies in a cell, and this bias varies between species. Protein expression can suffer if the CDS contains codons that are rarely used by the host cell - for example, due to a low concentration of the matching tRNA in the cytosol. Codon optimization tools are available which replace lower-frequency codons (based on the target organism) and minimize secondary structure that can impede translation - all without affecting the amino acid sequence.
3’ Untranslated Region (3’ UTR)
Description: Sequence between the stop codon and poly(A) tail.
Function in the cytosol: The 3’ UTR influences the stability of the transcript.
Considerations for therapies: As with the 5’ UTR, sequences from the alpha and beta globins are widely used for the 3’ UTR of therapeutic mRNAs. These sequences bind to proteins that enhance the transcript’s stability by deterring degradation of the poly(A) tail3.
Poly(A) Tail
Description: Polyadenylation at the 3’ end of the mRNA.
Function in the cytosol: The poly(A) tail regulates mRNA stability and promotes initiation of translation. It recruits poly(A) binding protein (PABP) which protects the transcript from 3’ to 5’ nuclease degradation. PABP also interacts with initiation factors at the 5’ end to trigger protein synthesis.
Considerations for therapies: Longer poly(A) tails generally confer greater stability to the transcript; the optimal length is 100-150 bases2. For in vitro mRNA preparation, a poly(A) tail can be generated by two methods:
- Enzymatic polyadenylation of the mRNA after in vitro transcription
- Incorporation of the poly(A) sequence in the DNA template prior to transcription
The latter method produces a more uniform length and enables better batch control of the product3. The downside is that poly(A) sequences greater than 120 bp can be unstable in plasmids: truncations during amplification in bacteria are common4,5. As a result, DNA templates may contain shorter and more heterogeneous poly(A) tails, which in turn makes mRNA products less stable and protein expression more variable. Using optimized DNA preparation protocols or segmenting poly(A) regions with short spacers5 can increase the stability of these sequences in plasmids.
For quality control, the poly(A) sequence should be verified by Sanger sequencing after final plasmid preparation to check for truncations and mixed populations. As the therapy advances in development and production is scaled up, full-length RNA sequencing can be used to thoroughly characterize the mRNA product, including the length distribution of poly(A) tails.
Pseudouridine Modification
Replacing uridine with pseudouridine (Ψ), an isomer with the same base-pairing properties, has been shown to reduce immunogenicity and increase the stability of mRNA therapies. Pseudouridine helps mRNA evade the immune system and increases protection against nucleases. It can be added to in vitro transcription reactions in form of ΨTP. Use of a modified pseudouridine contributed to the success of the mRNA COVID vaccines developed by Pfizer-BioNTech and Moderna Therapeutics6.
Conclusion
An ideal mRNA for therapeutic applications is optimized, from end to end, for efficient translation and enhanced stability. Once inside the cell, each mRNA component plays a critical role in producing strong protein expression.
Manufacturing well-designed mRNA in a fast, scalable, and reliable manner can be difficult if the entire production process is not robust or efficient. Read our tech note to learn how Azenta Life Sciences integrates gene synthesis and in vitro transcription to deliver high-quality mRNA. Also, we can help you design the 5’ cap, UTRs, and poly(A) tail of your mRNA construct as part of a pilot project.
References
1. Sikorski, P. J. et al. The identity and methylation status of the first transcribed nucleotide in eukaryotic mRNA 5′ cap modulates protein expression in living cells. Nucleic Acids Research vol. 48 1607–1626 (2020).
2. Kwon, S., Kwon, M., Im, S., Lee, K. & Lee, H. mRNA vaccines: the most recent clinical applications of synthetic mRNA. Archives of Pharmacal Research vol. 45 245–262 (2022).
3. Kwon, H. et al. Emergence of synthetic mRNA: In vitro synthesis of mRNA and its applications in regenerative medicine. Biomaterials vol. 156 172–193 (2018).
4. Grier, A. E. et al. pEVL: A Linear Plasmid for Generating mRNA IVT Templates With Extended Encoded Poly(A) Sequences. Molecular Therapy - Nucleic Acids vol. 5 e306 (2016).
5. Trepotec, Z., Geiger, J., Plank, C., Aneja, M. K. & Rudolph, C. Segmented poly(A) tails significantly reduce recombination of plasmid DNA without affecting mRNA translation efficiency or half-life. RNA vol. 25 507–518 (2019).
6. Morais, P., Adachi, H. & Yu, Y.-T. The Critical Contribution of Pseudouridine to mRNA COVID-19 Vaccines. Frontiers in Cell and Developmental Biology vol. 9 (2021).